CRISPR's path from curiosity to Nobel Prize (1987-2020)
1987-2020What Happened
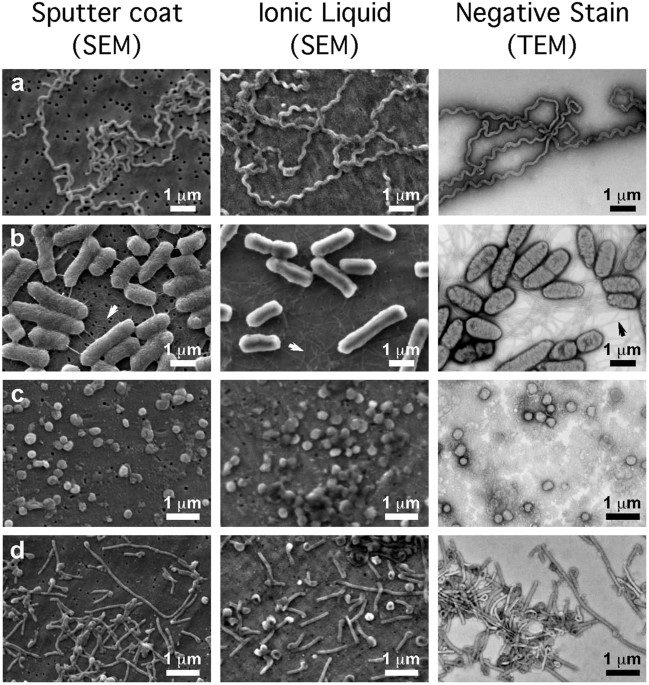
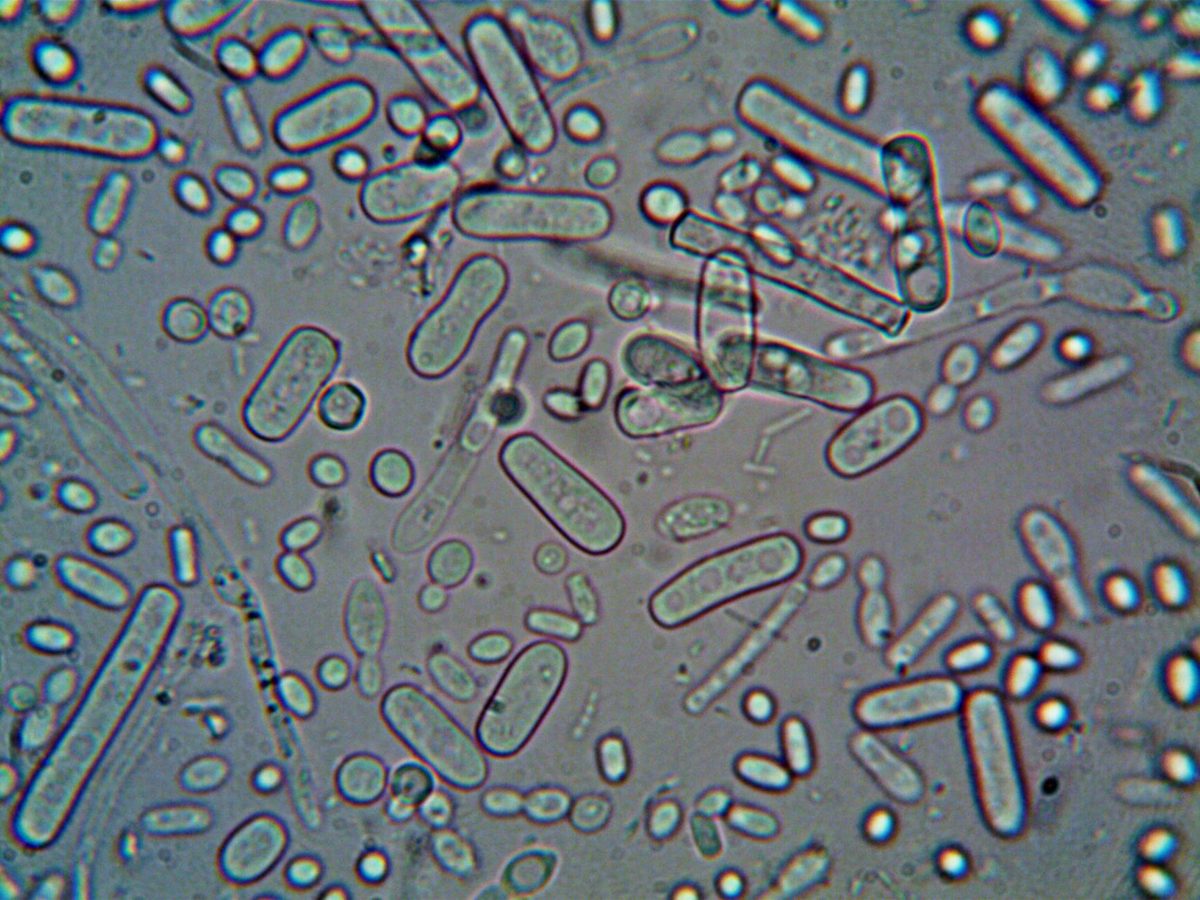
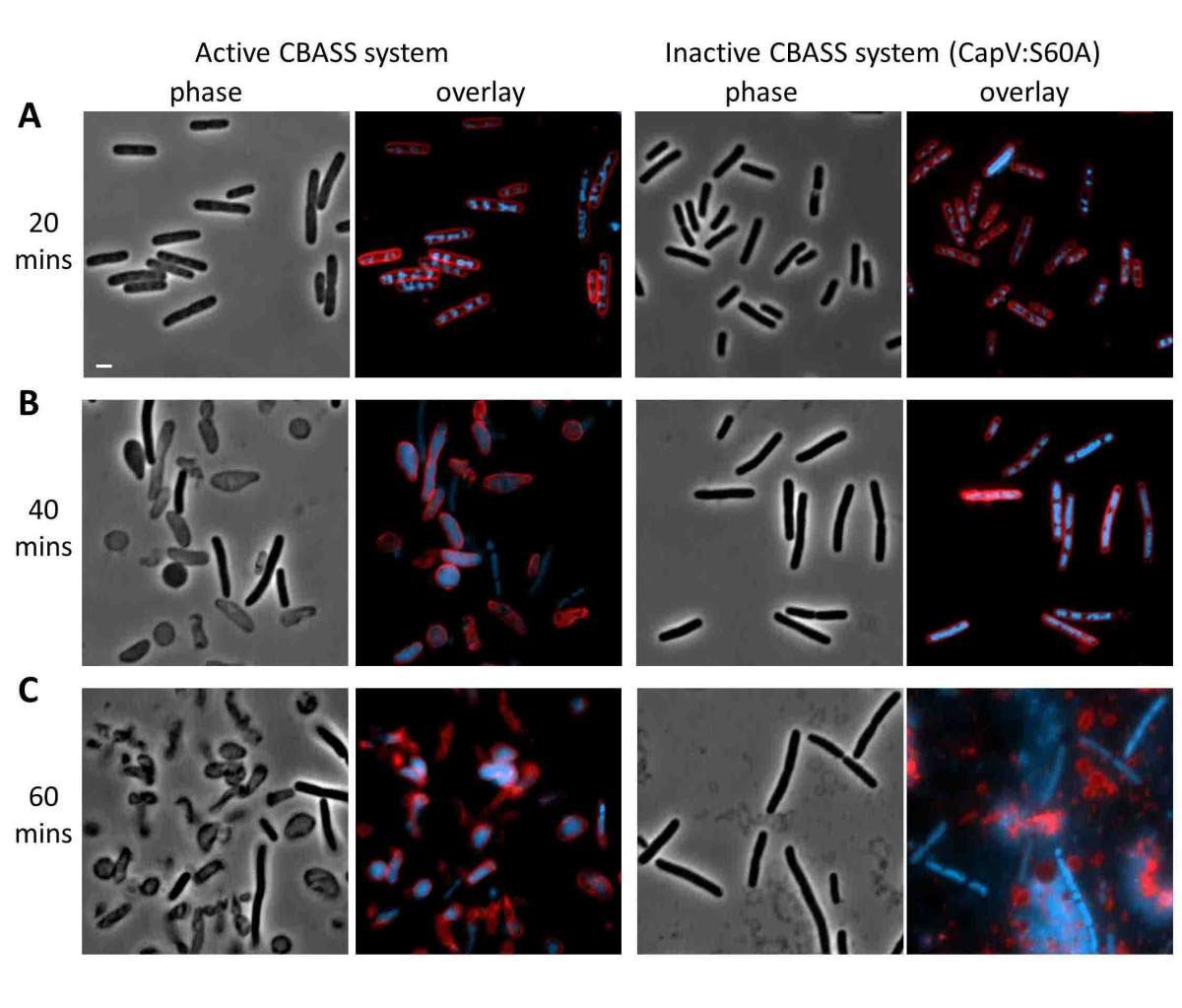
Repeat sequences noticed in E. coli in 1987 sat unexplained for 18 years. Mojica proposed in 2005 that they were a bacterial immune system; Barrangou and Horvath proved it in 2007 using yogurt cultures; Doudna and Charpentier turned it into a programmable gene-editing tool in 2012. The 2020 Chemistry Nobel followed.
Outcome
Gene editing became routine in research labs within three years of the 2012 paper.
CRISPR-based therapies for sickle cell disease received FDA approval in 2023; the technology underpins a multi-billion-dollar biotech sector.
Why It's Relevant Today
Every bacterial defense system discovered since 2018 is being evaluated for the same trajectory. Clover's programmable nucleotide-switch logic is exactly the kind of feature that made CRISPR valuable.